MIATool
Microscopy image analysis
The Microscopy Image Analysis Tool (MIATool) is a software application designed for the viewing and processing of large sets of image data. It makes efficient use of memory by working with pointers to images rather than the images themselves, and it saves disk space by storing reusable processing parameters in lieu of actual processed images on the hard drive. In addition, MIATool allows for multiple interpretations of a given image data set, realized by different, in general N-dimensional, arrangements of pointers to the images. A physical image data set can therefore be represented, viewed, and processed independently in different ways.
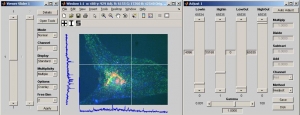
MIATool supports the viewing and processing of an N-dimensional array of images. At its core is an image viewer which allows the traversal of an N-dimensional array of images. Besides the standard display as pixels of varying intensity values, options are available to view the images as mesh or contour plots. In addition, options exist that allow the concurrent viewing of multiple images, either in parallel in separate windows, or as an RGB overlay. Useful features such as the reporting of pixel intensity values, the optional display of a crosshair, and the display of pixel intensity profiles along the row and column of the crosshair, are also supported.
The current version of MIATool supports four different image editing tools which can be used to process the images displayed in the viewer. The intensity adjustment tool provides different ways to modify the pixel intensity values, and the crop tool allows trimming of the images to retain only the portion that is of interest. The two remaining tools – the segmentation tool and the label tool – can be used for manual image segmentation and image labeling. For more details concerning the design of MIATool , please refer to [1].
References
[1] Chao, J., Ward, E. S., and Ober, R. J. (2010). A software framework for the analysis of complex microscopy image data. IEEE Transactions on Information Technology in Biomedicine, 14: 1075-1087, 2010.
[2] Chao, J., Long, P., Ward, E. S., and Ober, R. J. (2007). Design and application of the Microscopy Image Analysis Tool. Engineering in Medicine and Biology Workshop, 2007 IEEE Dallas, pages 94–97.
Currently Available Version
V1.1 (May 18, 2009) Please note that this is a Beta version.
Minimum System Requirements
- PC with Pentium IV or equivalent processor suggested
- Windows XP Service Pack 2 *
- 512 MB RAM
* MIATool was implemented and tested on Windows XP Service Pack 3. However, the executable provided may run on other versions of the Windows operating system as well.
Documentation
Download MIATool User’s Manual 
(contains installation instructions and step-by-step tutorial)
How to obtain the software
If you are interested in access to one of our software, please contact us at info@wardoberlab.com
FAQ
None available as yet.
Release Notes
- More variants of the 16-bit tiff format supported for input images
- 8-bit tiff format support for input images.
- As a trade-off with better support for the tiff format, image creation, loading, and viewing are slower than in V1.0
- BSD licensed
Bug Reporting
We are constantly trying to improve this software and would be very grateful if you could send us a report on any bugs you find. Please send all bug reports to the email addresssoftware@wardoberlab.com with the subject MIATool Bug. Please include as many details about the bug as you can, including details of all tasks that led up to exposure of the bug. This will help us in finding a solution to the problem. When we have a fix for the problem, we will release an updated version of the software. Please look in the Release Notes section for problems we are already aware of and have solutions for.
MIATool Development Team
Current Members
- Raimund J. Ober (Team Leader)
- Jerry Chao
- Anish V. Abraham
- E. Sally Ward
Past Members
- Sripad Ram
- Jincheng Pang
- Hongguang Xi
- Xuming Lai
- Palmer Long
- Piyush Gehalot
- Lalit Raina
- Prashant Prabhat
Acknowledgments
The development work for MIATool was supported in part by the National Institutes of Health.
Contact Address
Raimund J. Ober, Ph.D.Professor |
E. Sally Ward, Ph.D.Professor |
